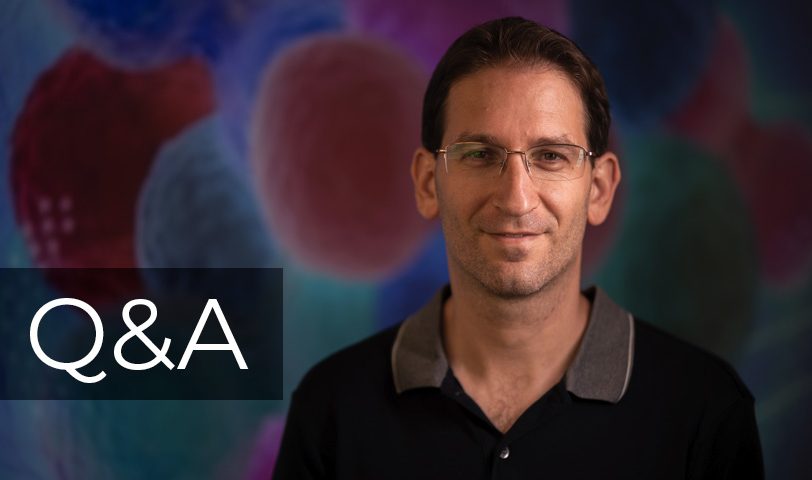
Dr. Itay Tirosh is a member of the faculty of the Weizmann Institute of Science in Molecular Cell Biology. He returned to Weizmann, where he did his PhD in the Department of Molecular Genetics, after completing a postdoctoral fellowship at the Broad Institute of MIT and Harvard. There, he pioneered the application of single-cell RNASeq to solid human tumors and established a general approach to studying intra-tumor heterogeneity in collaboration with clinicians at Boston hospitals, demonstrating how single-cell genomics offers insights with implications for both targeted and immune therapies.
His joint computational and experimental research group studies solid tumors as a complex ecosystem, and find potential improvements in cancer research and cancer therapy.
Please describe your current research, the focus of your lab, and the practical implications of your research
The lab combines computational and experimental methods to study human tumors as a complex ecosystem in which diverse cancer and non-cancer cells interact and collectively determine tumor biology and response to therapies.
Every tissue is very complex and contains many different cells. In the past it was very difficult to understand those differences, but with single cell sequencing we are able to study them directly, especially in the context of tumors. Beyond understanding the tissue composition, we can try to examine the main reasons why we can’t cure cancer. There are two main problems. When we treat tumors, we kill off many cells but for a large percentage of people who are told they are cancer-free, hidden cancer cells are left behind. Tumors come back from the remnant of the original tumor, dormant cells that regrow into a new tumor. Why these cells are not eliminated by the treatment, even though we know some cells differ from one another, is what we are trying to determine.
The second problem is metastasis. How does this happen? These cells are part of a mass – somehow some are capable of detaching from the tumor, invading other tissues and adapting to those tissues, making it more difficult to cure. This relates back to the diversity of cells and defines the main bottlenecks in finding a cure.

We want to understand this diversity in order to eliminate these unique cells – either the ones that evade treatment or cause metastasis.
We work closely with clinicians to get patient samples directly and break them down into individual cells and then measure the activity of genes in each of those cells. This provides us with a lot of data: what genes are expressed in each cell, or in other words, the fingerprint of each cell. For each cell we want to know which genes are active and based on that to understand which cells are able to do the above two things – evade treatments and metastasize. The main cancers we look at are brain cancers, primarily glioblastomas, and head and neck cancers.
Even though tumors are unique in each patient, we see that differences between cells have a very typical patterns. For example, two patients with glioblastomas have very different tumors, but the internal structure within each of them is very similar. It’s like comparing two rooms that are very different but both have tables and chairs. We focus on the patterns, or groups of cells, that are relatively similar. Our initial observation is that the way in which they evade treatment is probably similar as well.
What do you enjoy most about your research?
I like the multidisciplinary approach and atmosphere of the lab. Even though we are ultimately interested in cancer, some lab members are biologists, others are physicians and most have a computational background and focus on analyzing data. We take a system like a tumor, profile it, and generate a ton of data – millions of pieces. The real challenge is making sense of the data, so most of the time revolves around computational analysis and biological interpretations of the results.

What inspired you to pursue this area of research?
I came from systems biology where we want to understand every biological component and how they interact in a complex system, and I wanted to apply this approach to cancer. The single cell methods were used a lotat the Broad Institute of Harvard/MIT where I did my postdoc and we were the first to apply this to tumors.
What does it mean to you to be part of the Zuckerman Faculty Scholars Program?
Every lab needs support to get started. The Zuckerman Program enabled me to establish my lab, purchase equipment, and get started with my research. Beyond that, it’s been very helpful to be part of such an impressive community. As a Zuckerman Scholar I continue to collaborate closely with scientists in the United States and am now taking a sabbatical there.
Where do you hope your research will have the greatest impact?
I hope that my research will go beyond the state it is in now to have an actual impact on cancer treatment. We identify important tumor subpopulations such as cancer stem cells, drug resistant cells, invasive cells, and immune cells that respond to immunotherapies. We then study their function, regulation, and vulnerabilities, and our ultimate, long-term goal is to develop better cancer treatments. Working closely with clinicians could help to make the transition from basic discoveries to clinical trials and ultimately having a clinical impact.

 ISRAELI COUNCIL FOR HIGHER EDUCATION
ISRAELI COUNCIL FOR HIGHER EDUCATION MIT-Israel Zuckerman STEM Fund for Faculty Collaboration
MIT-Israel Zuckerman STEM Fund for Faculty Collaboration The Zuckerman Travel and Research STEM Fund at Harvard
The Zuckerman Travel and Research STEM Fund at Harvard Zuckerman AI Fund at Technion
Zuckerman AI Fund at Technion Alan Alda Communicating Science
Alan Alda Communicating Science Zuckerman Institute – ScienceAbroad
Zuckerman Institute – ScienceAbroad Zuckerman Institute – America-Israel Friendship League partnership
Zuckerman Institute – America-Israel Friendship League partnership